REPLi: Identify Protease Substrates
Do you have an unidentified protease? Do you want to know its target sequence?
A Rapid Endopeptidase Profiling Library (REPLi) is a generic Fluorescent Resonance Energy Transfer (FRET) Peptide Library designed specifically to overcome common hurdles and simplify the protease profiling process. Mimotopes’ REPLi Library is available off-the-shelf and ready to use from our online catalogue.
Find out more
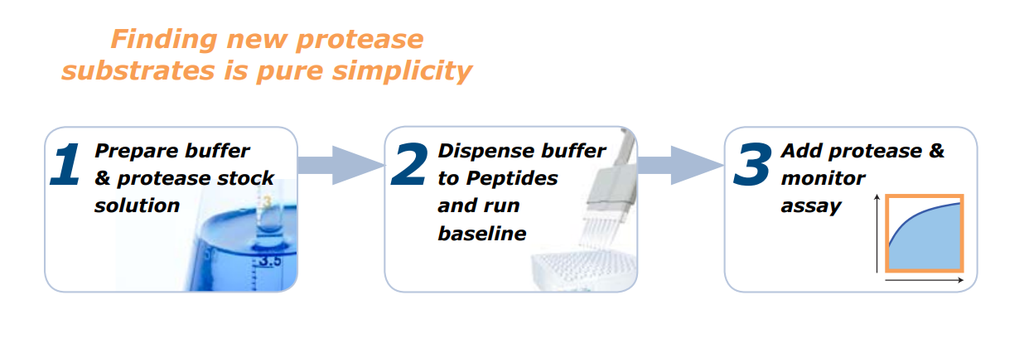
REPLi Requirements
An advantage of REPLi is it doesn’t require any prior knowledge of the proteases under investigation to help define the sequence of the protease cleavage site.
This peptide library comprises 3375 peptides divided into 512 distinct pools, each containing only 8 peptides so it can be rapidly deconvoluted to identify the specific sequences cleaved by your proteases.
Proteases are valuable drug targets in disease treatment, therefore fast identification of protease substrates directly from cells and tissues is invaluable.
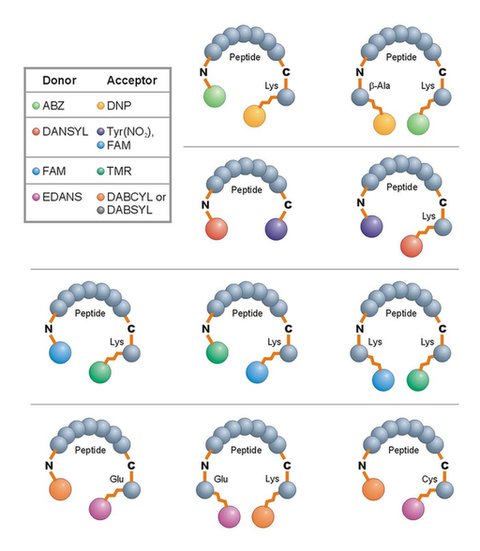
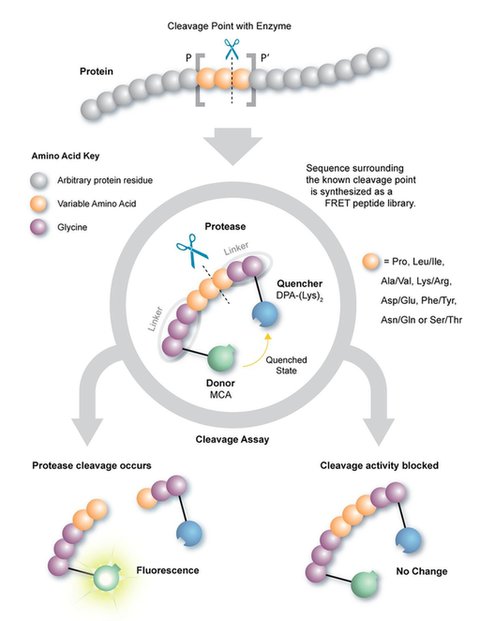
How FRET Functions
FRET is commonly used to investigate protein interactions, including proteolysis.
In REPLi, the tripeptide core sequence is book-ended by glycine residues, with a fluorescent donor and quenching acceptor on opposite ends. When proteases cleave the peptide sequence, the quenching effect ends and the donor fluoresces.
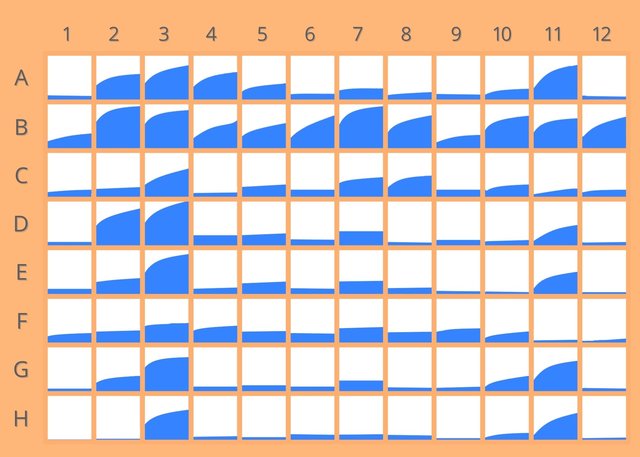
Through real time proteolytic activity monitoring, this peptide library design provides a quantitative measure of the differences in substrate processing of low-abundant proteins, as disease markers.
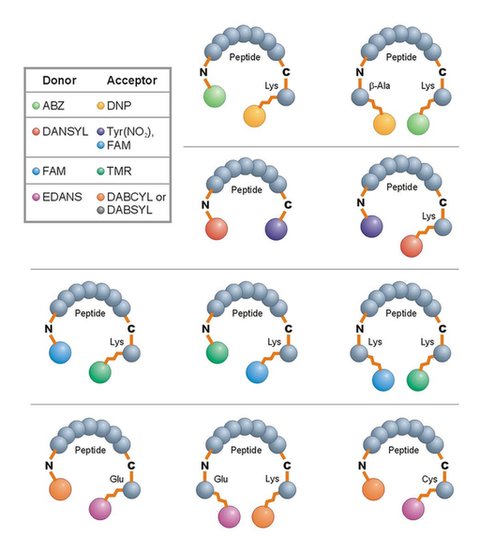
Is REPLi Reliable?
Validation studies with well-characterised representative proteases from each of the four mechanistic protease classes have shown the impressively specific and sensitive capabilities of REPLi.
Results obtained from REPLi can be followed up with precise directed analysis using purified custom peptides or peptide libraries.

CONTACT US TODAY
for a peptide quote!
Australia
North and South
America
asia
UK / Europe / Africa / Middle East
☎ +61 3 9565 1111 |
☎ +1 651 603 0909 |
☎ +61 3 9565 1111 |
☎ +44 (0)151 556 0547 |

Contact us today
for a peptide quote!
Australia
☎ +61 3 9565 1111 |
North and South America
☎ +1 651 603 0909 |
asia
☎ +61 3 9565 1111 |
UK / Europe / Africa / Middle East
☎ +44 (0)151 556 0547 |

Contact us today
for a peptide quote!
Australia
☎ +61 3 9565 1111
North and South America
☎ +1 651 603 0909
asia
☎ +61 3 9565 1111
UK/ Europe/ Africa/ Middle East
☎ +44 (0)151 556 0547


