Insight
Why the German Centre for Diabetes Research changed its approach to handling data
How can life science researchers study large datasets and uncover potential new insights with the power of graph databases. CEO of graph database company Neo4j Emil Eifrem has plans to use this talent to change the way we discover new drugs.
The emergence of big data and other advances in technological innovation has provided medical researchers, among others, with the opportunity to attain valuable, previously unobtainable insight with the potential to improve all of our lives. The problem: real-world data comes in multiple, highly unstructured formats.
Why is that a problem? To really start exploiting the power of things like big data, researchers must go well beyond what they’re doing now – simplistic managing, analysis and storage of data. This means taking a cold and objective look at the tools they’ve historically used in research, especially spreadsheets and relational database technology.
That’s because traditional methods can’t cope with the volume, as well as the inconsistent nature, of the kind of data we want to study in-depth now. By its nature, medical data is heterogeneous: it can run from cell-level to detailed data to macro-scale disease network tracking across different diseases areas, with scientists often wanting to link both ends of that spectrum. Realistically, the breakthroughs we want to find are likely to come from working with many different data sets, as that is where the interesting results can lie, but it is a real challenge to model this.
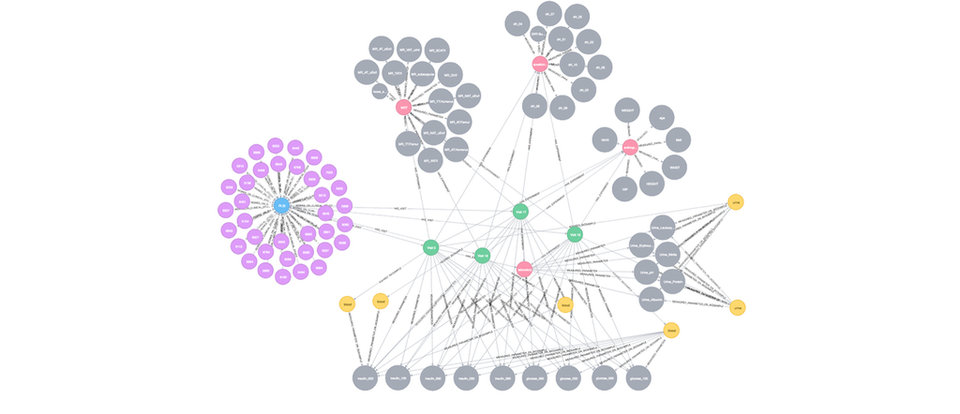
Graph representation of meta data for DZD’s 'Prediabetes Lifestyle Intervention Study'.
Image courtesy of DZD
Utilising graph technology
Another obstacle has been the fact that it can be really hard to spot relationships between things in traditional computerised ways of storing information. That’s where something called graph technology is starting to be evaluated, as it is all about those relationships. Graph software hit the world stage when it emerged that it was the technology behind the world’s largest investigative journalism project, the Panama Papers, where a team of global researchers went through a massive three terabyte data set spotting connections that would have been an impossible task using traditional data technology approaches.
It turns out that large, complex datasets like those researched in the Panama (and its successor, the Paradise) Papers are too difficult for the traditional business database technology we use to work with data today, the relational database, to easily work with. This is down to the underlying fact that relational databases model the world using sets of tables and columns carrying out complex joins and self-joins to perform queries. But, the questions that need to be asked in areas such as Life Sciences are quite hard to construct and expensive to run in this form, while making them work in real time is a big ask, with performance degrading as data volume grows.
Large, complex datasets are too difficult for the traditional business database technology we use to work with data today.
Also humans look to model connections between data elements in a very different way, and the relational data model doesn’t really line up at all with our mental visualisation of problems, something technically defined as “object-relational impedance mismatch”. Trying to take a data model based on relationships and pushing it into a tabular framework, the same way a relational database does, ultimately creates disconnect that can not only cost valuable time, but can also lead to missing potentially useful patterns and leads.
This is where graph software can step in, however – as its ‘unique selling point’ is its innate ability to discover relationships between data points and understanding them, and at scale. That is why it is an exciting tool for medical researchers, as it’s starting to enable them to uncover hidden patterns when delving into the difficult problems, with emerging use cases ranging from new molecule research to clinical trial work.
Posing useful questions in natural language
A great example is the German Centre for Diabetes Research (DZD). Based in Munich, the DZD is a national association that brings together experts in the field of diabetes research and combines basic research, translational research, epidemiology, and clinical applications.
The idea is to find treatment measures for diabetes across multiple disciplines and to see what treatment the latest biomedical technologies may offer the growing number of citizens dealing with the condition: in order to better understand diabetes’ causes, its scientists examine the disease from as many different angles as they can. DZD’s research network thus accumulates a huge amount of data that is in turn distributed across various locations – and its internal IT leadership decided it needed a better way of seeing all this in the round.
It’s done that by building a ‘master database’ to consolidate all this information and provide a better way of visualising it so as to provide its 400-strong team of scientist peers with a properly holistic view of available information, enabling them to gain valuable insights into the causes and progression of diabetes.
When the centre’s head of bioninformatics and data management, Dr Alexander Jarasch, was looking for the right data tool to build such a system, he drew on experience gleaned from previous work on a project at Munich’s Helmholtz Zentrum, where he’d tried to link DNA sequence data with metabolomics data (data for the systematic capture of metabolic matter). That had proven a difficult area to make any progress on until he and his team discovered a graph database approach – a positive experience that prompted him to test graph technology at DZD, specifically software from the native graph database, Neo4j.
DZD’s research network accumulates a huge amount of data that is in turn distributed across various locations.
As a result, Jarasch has built a new internal tool, DZDconnect, built in Neo4j that sits as a layer over the centre’s various relational databases, finally joining up different DZD systems and data silos. When fully implemented the expectation at the centre is that, using DZDconnect, its teams can access metadata from clinical studies and exploit the visualisation and the easy querying it has made possible.
The promise here is that the more detailed the information, the easier it is to identify relationships and patterns. The idea is to eventually enable DZD’s researchers to pose useful questions in a Natural Language form, such as: ‘How many blood samples have we received from male patients under 69? Which studies are the samples from? Which parameters got measured this time?Over what period of time was the sugar value measured? Were measurements taken on an empty stomach or after a glucose intake? How has the value changed in the long term? Can the change be attributed to a healthier diet, a drug or a hitherto unknown factor?’
Jarasch confirms that, “With graph we were able to combine and query data across various locations. Even though only part of the data has been integrated, queries have already shown interesting connections which will now be further researched by our scientists”. In the long term, as much DZD data as possible should be integrated into graph database, Jarasch believes. The next step is to see how human data from clinical research will be complemented with highly standardised data from animal models, such as mice, to find communalities.
Interestingly, it’s not just graph technology being employed in this new way of working with complex data at DZD. Artificial intelligence (AI) techniques like Machine Learning will play a key role going forward, says DZD, with a particular area of interest building a system able to ‘read’ scientific texts and integrate them into the database ready for analysis. "Technology makes it easier to view medical issues from different perspectives and across indications," he points out. "This also makes it possible to identify correlations between various common diseases."
Making some useful unknowns ‘known’
The kind of innovative data management and analysis approach DZD is pioneering in Munich could well be the way forward in precision medicine, prevention and treatment of diabetes – and, perhaps, for other diseases. There are at least eight significant cancer research projects we know of that are using graph databases, for instance.
Graph technology’s innate ability to discover relationships between data points allows us to understand them.
Graph technology’s innate ability to discover relationships between data points allows us to understand them, which could have an enormous role to play in medicine and healthcare in the future. And there’s also the promise that making these data connections is the beginning of a journey that could lead to real breakthroughs that older ways of working with data just can’t offer. Further input from developers to build graph-based data structures for research will enable ever more highly trained specialists to have access to data in a form they can work with much earlier in their research.
To sum up, in data everything is connected, but sometimes the connections are hard to spot with the technology we’ve been using so far. Having the power to make the unknowns known, uncovering hidden promise, could be a compelling tool in life science research – and one that is already having an enormous impact on the industry.
Go to top
